New tools to manipulate biology
Chemistry has provided many key tools and techniques to the biological community in the last twenty years. We can now make proteins that Mother Nature never thought of, image unique parts of live cells and even see cells in live animals. In ACS Central Science, three independent research groups from the University of Geneva (UNIGE) and one from the University of Basel (UNIBAS) take these accomplishments a step further, reporting advances in both how proteins are made and how you can see their expression patterns in live animals.
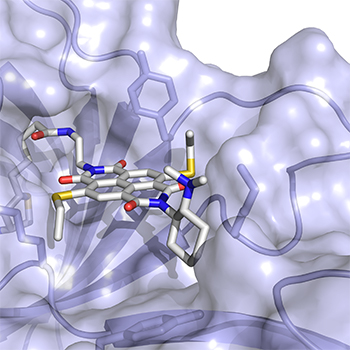
Proteins are the workhorses of every cell. They are made up of building blocks called amino acids that are linked up and fold together into functional machines to power every major cellular process. To do these tasks, nature relies on twenty of these blocks together with a few special “co-factors,” often vitamins. However, chemists have discovered clever ways of expanding a protein’s repertoire, engineering in different amino acids or co-factors than you would find in natural biology. Stefan Matile, Thomas Ward and coworkers designed a new co-factor that reverses a classic protein interaction called the cation-Π, meaning the stabilization of a positive charge on an electron-rich molecular plane. Nature uses this cation-Π interactions to prepare molecules as important as steroids, hormones, vitamins, visual pigments or fragrances, to transduce signals in the brain, to recognize antigens, and so on. Using their new co-factor, and resulting artificial protein, Matile and Ward’s groups collaborated to create the first “anion-Π ” enzyme where that electron-rich molecular plane is replaced by an electron-poor plane to stabilize a negative rather than a positive charge during a molecular transformation. In a test tube, proteins with this new-to-nature functionality were able to outperform traditional organic catalysts in an important but disfavored addition reaction with high specificity and selectivity. They believe their approach can be moved to work in cells and can help make other currently impossible chemical transformations a reality.
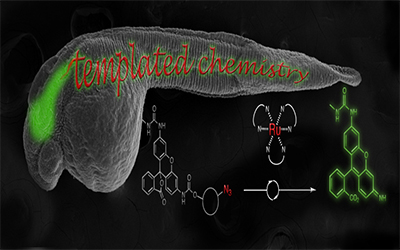
Meanwhile, Nicolas Winssinger and his lab recognized the opportunity to use chemical reactions to visualize mRNA in live animals. All proteins come from mRNA, so while DNA is the blueprint of life, mRNAs provide the work-orders which are further regulated by microRNAs. Being able to see these can tell you a lot about what is happening to cells and animals in real time. However, there are few tools that currently enable you to see the RNA in live animals, and those that exist usually rely on complicated genetic strategies. Winssinger and Gonzalez-Gaitan’s teams designed a templated-reaction that yields fluorescent markers to specific RNAs of interest. Their chemistry is so mild that it works in live zebrafish embryos without disturbing them, and because it does not require genetically modified organisms, it could be quickly adapted for other systems. Finally, they posit that their method could be expanded towards theragnostics, where you could simultaneously see and target treatments to the location of disease’ molecular basis.
Contact: Stefan Matile +41 22 379 65 19 Nicolas Winssinger +41 22 379 35 63
24 May 2016