HDX-MS

HDX-MS methodology :
The methods allows you to monitor protein dynamics based on protein amide proton exchange with the solvent. Starting from a purified sample, dynamics can be monitored for an isolated protein and for the same protein is different conformations, for example when bound to a ligand. The ligand can be anything like another protein, small molecule, nucleic acid of membranes (lipid vesicles, nanodiscs and monolayers). Comparison of the dynamics in absence and presence of the ligand will identify both i) ligand binding sites as well as ii) regions of the protein allosterically modified upon ligand binding.

HDX-MS sample preparation
Concentrated protein sample is diluted in deuterated buffer for a set amount of time. Following deuteration, Hydrogen/Deuterium exchange is quenched by reducing pH to 2.5 and temperature to 0 °C. In parallel, protein is denatured and the sample is flash frozen in liquid nitrogen where it can be stored up to four weeks.
For deuterium quantification, sample is digested online with pepsin (at 0 °C and pH 2.5), peptides are desalted and their mass are analyzed by Mass Spectrometry
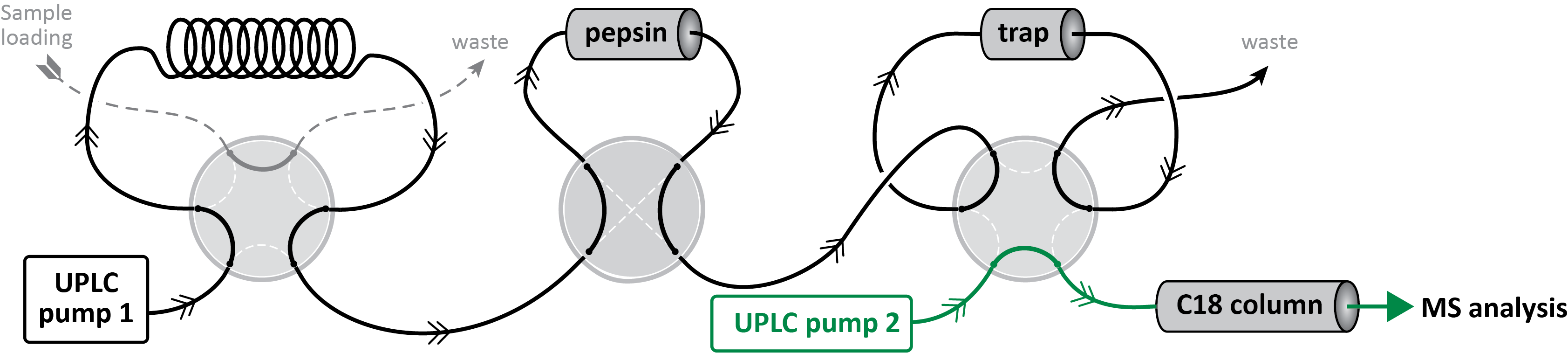
Instrumentall workflow
Following manual injection into the loop, sample is digested by passing through a pepsin column and the peptides are trapped into a C18 reverse phase column. After washing, a second pump applies a water to acetonitrile gradient to detach the peptides from the trap, separate them on a 10 cm C18 reverse phase column prior to mass analysis using an orbitrap instrument
Sample requirements :
To run a HDX-MS experiment, we will need both the isolated recombinant protein target as well as the purified protein-ligand complex. Alternatively, if the protein-ligand complex cannot be isolated, we can also work with an excess of the ligand to obtain maximal occupancy on the ligand-binding site. The amount of protein needed depends on the MS signal and this can only be determined by doing a "feasibility assay".
For a feasibility test we will need 150 microgram of target protein and of the complex
For a "full" experiment we expect to need a total of 550pmol of each: i) the isolated recombinant ligand and ii) the complex. When working with an excess of the ligand, we will need larger amount of the ligand over the target protein. A coomassie-stained gel of each studied protein is requested to have an estimate of protein purity.
Buffer and protein details :
Proteins need to be in a defined buffer without detergent. Ideally all samples (proteins, ligands, lipids, nucleic acids, ...) should be in the same buffer.
Protein concentrations should be as high as possible, ideally in the 1-4 mg/ml range.
The exact sequence of the studied protein, including tags, must be provided before the start of the experiment
Experiment design 1 – feasibility test :
Before starting the experiment a feasibility test will be run that is essential to determine whether the recombinant protein can be studied by HDX-MS. Following parameters will be evaluated and optimized :
- Amount of protein necessary to have a good MS signal for each run
- Protein coverage and peptide redundancy
- Best quenching conditions
If the listed parameters are fine (approximately 95% of cases), then we proceed with a first deuteration experiment at a single deuteration timepoint to monitor the overall dynamics of the protein.
Experiment design 2 – full experiment :
To identify binding site of the ligand on the recombinant protein, we will monitor the H/D exchange rate of two conditions: 1) recombinant protein alone vs 2) recombinant protein in complex with ligand. For each of the two conditions, we will monitor H/D exchange rates for three different timepoints (3sec, 30sec and 5min) to capture both the fast-exchanging amide hydrogen exchange and slow-eschanging ones. Each of the reactions need to be run in triplicate for publication.
Comment: deuteration reactions can be done at room temperature or at 0 °C. This needs to be discussed for each individual project.
Suggested references :
Reviews on HDX-MS methodology
James EL. et al, Chem Rev 2022 DOI: 10.1021/acs.chemrev.1c00279
Krishnamurthy S et al Curr Opin Struct Biol 2025 DOI: 10.1016/j.sbi.2025.103051
Engen JR. et al, Trends Biochem Sci 2020 DOI: 10.1016/j.tibs.2020.05.005
Trabjerg E et al, Trends in Analytical Chemistry 2018 DOI: 10.1016/j.trac.2018.06.008
Masson GR et al, Expert Opin Drug Discov 2017 DOI: 10.1080/17460441.2017.1363734
Chalmers MJ et al, Expert Rev Proteomics 2011 DOI: 10.1586/epr.10.109
Conformational changes observed by HDX-MS : protection vs exposure
Konermann L. et al, Analyst 2014 DOI: 10.1039/c4an01307g
Inspiring publications for figure generation
Gu W. et al, Nat Commun 2021 DOI: 10.1038/s41467-021-24929-5
David SC et al, mSphere 2023 DOI: 10.1128/msphere.00226-23
Calugareanu D et al, J Mol Biol 2022 DOI: 10.1016/j.jmb.2021.167356
Zhang N et al, Nucleic Acid Res 2022 DOI: 10.1093/nar/gkac170
Jia R et al, Nat Commun 2020 DOI: 10.1038/s41467-020-20032-3
Garcia NK et al, Structure 2015 DOI: 10.1016/j.str.2015.02.006
Vadas O et al, Proc Natl Acad Sci USA 2013 DOI: 10.1073/pnas.1304801110
HDX-MS of integral membrane proteins and membrane-interacting proteins
Javed W et al, Essays Biochem 2023 DOI: 10.1042/EBC20220173
Stariha JTB et al, Methods Mol Biol 2021 DOI: 10.1007/978-1-0716-1197-5_22
Example with EX1 kinetics (bimodal distribution)
Karki R. et al, Biochemistry 2023 DOI: 10.1021/acs.biochem.3c00338
Merkle PS et al, Sci Adv 2018 DOI: 10.1126/sciadv.aar6179
Tuttle LM et al, J Am Soc Mass Spectrom 2025 DOI: 10.1021/jasms.4c00471
Reference paper on best practice in HDX-MS
Masson GR et al, Nat Methods 2019 DOI: 10.1038/s41592-019-0459-y
Softwares for data analysis and visualisation
Other groups using HDX-MS to tackle biological questions
Burke Lab | University of Victoria, Canada
Englander Lab | University of Pennsylvania
Engen Lab | Northeastern University
Guttman Lab | University of Washington
Konerman Lab | University of Western Ontario, Canada
Masson Lab | University of Dundee, UK
Politis Lab | University of Manchester
Rand Lab | University of Copenhagen
Schriemer Lab | University of Calgary, Canada
Williams Lab | MRC-LMB Cambridge, UK
