BioRxiv, 2026
Our results associate alterations in Puf3 post-translational status and function, including contribution to translation elongation dynamics, with not4Δ mutant phenotypes.
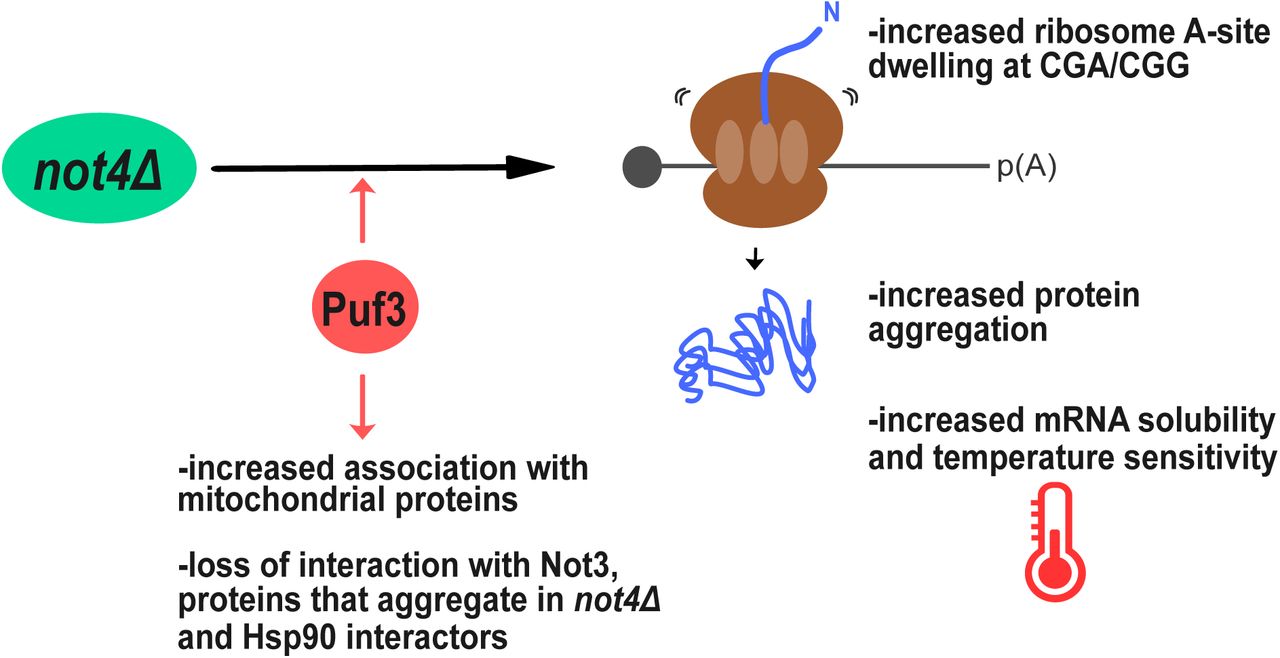
Puf3 contributes to changes in mRNA solubility, translation elongation dynamics at rare arginine codons and loss of protein homeostasis in cells lacking Not4
Abstract
The Not proteins of the Ccr4-Not complex regulate translation elongation dynamics, essential for proper folding and assembly of new proteins. In yeast, ribosomes with non-optimal codons in the A-site are enriched within the pool of ribosomes bound by Not4 and Not5. Such ribosomes accumulate in cells lacking Not4 or Not5 that show defects in co-translational assembly and aggregation of new proteins. Recently we observed that depletion of Not1 and Not4 inversely regulate changes in mRNA solubility, correlating with inverse codon-specific changes in A-site ribosome dwelling occupancies (RDOs). Here we describe that mRNAs less soluble upon Not4 depletion are enriched for targets of the RNA-binding protein Puf3. We determine that Puf3 contributes to inverse changes of A-site RDOs upon Not1 and Not4 depletion, in particular at rare arginine codons, and it contributes to changes in mRNA solubility in not4Δ. Moreover, deletion of Puf3 suppresses temperature sensitivity and protein aggregation in the not4Δ strain, while overexpression of Puf3 is toxic. Puf3 post-translational modifications and the Puf3 interactome are altered in not4Δ. Taken together, our results associate alterations in Puf3 post-translational status and function, including contribution to translation elongation dynamics, with not4Δ mutant phenotypes.
